Operations and Indexing
Concatenation
You concatenate vectors using the c function; this is the standard way to
construct vector objects:
x <- c(1.2, 3.8, NA, 4.4, 1, 2.718)
y <- c(8, 9)
z <- c(x, y)Recall that there are no scalars in R, so 1.2 is a length-1 vector. The
c function creates a new vector by concatenating its arguments together.
Repeating
The rep function constructs a new vector by repeating the elements of
its argument.
rep(y, 3)[1] 8 9 8 9 8 9
rep(y, each = 3)[1] 8 8 8 9 9 9
The first call repeats the input vector y three times; the second call
repeats each element of y three times.
Vector Operations
Most functions in R operate on vectors, and they do so in an element-wise fashion:
log(x)[1] 0.1823 1.3350 NA 1.4816 0.0000 0.9999
x^2[1] 1.440 14.440 NA 19.360 1.000 7.388
Binary operators take corresponding elements from their arguments:
a <- c(1, 0, -2, NA, 10)
b <- c(2, NA, 3, 7, 4)
a + b[1] 3 NA 1 NA 14
a * b[1] 2 NA -6 NA 40
Note that if either argument is NA, then the result is NA.
Comparison operations result in logical vectors:
a < b # less than[1] TRUE NA TRUE NA FALSE
a == b # equal to (NOTE double equals)[1] FALSE NA FALSE NA FALSE
a != b # not equal to[1] TRUE NA TRUE NA TRUE
Other valid comparison operators include <=, >, and >=.
Application: Transforming Variables
Vector operations are useful for transforming variables. For example, in the original bike data, passing distance is slightly skewed to the right:
bikedata <- read.csv("bikedata.csv")
hist(bikedata$passing.distance, col = "firebrick")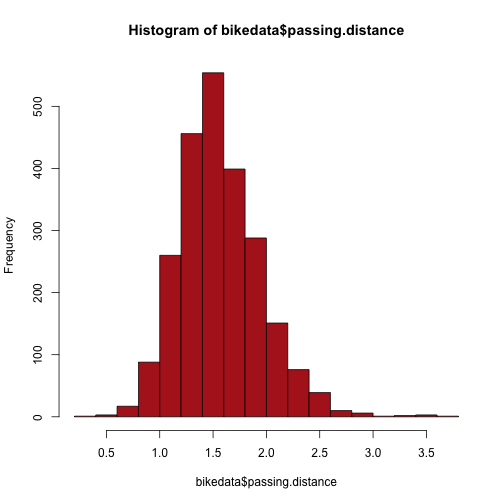
We can adjust for some of this skewness by working with the square root of passing distance:
hist(sqrt(bikedata$passing.distance), col = "steelblue", main = "Square Roots are more symmetric")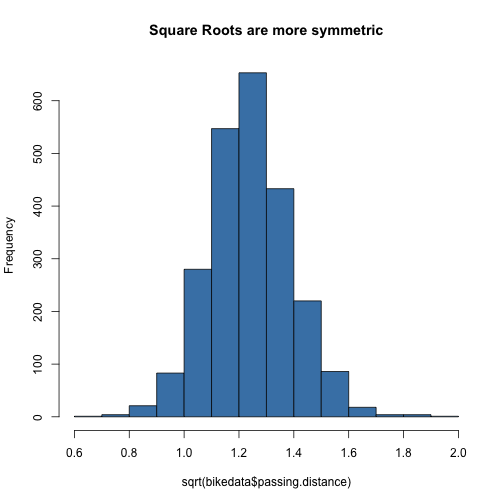
Recycling
When the arguments to a vector function have unequal lengths, the elements of the shorter argument get recycled. This is useful for cases when one argument is a element vector:
10 - x[1] 8.800 6.200 NA 5.600 9.000 7.282
x/2[1] 0.600 1.900 NA 2.200 0.500 1.359
Logically, the above two operations are equivalent to
rep(10, length(x)) - x and x / rep(2, length(x)).
The recycled vector can have arbitrary length, as in the example
c(1, 2, 3, 4, 5, 6) * c(-1, 1)[1] -1 2 -3 4 -5 6
Recycling also works in cases where the shorter vector’s length does not divide the longer vector’s, but this is considered bad programming style, and R will issue a warning in such situations.
c(1, 2, 3, 4, 5, 6, 7) * c(-1, 1)Warning: longer object length is not a multiple of shorter object length
[1] -1 2 -3 4 -5 6 -7
Application: Testing the Empirical Rule
Let’s using recycling and the operations we have learned so far to check if the Empirical Rule holds for the square root of passing distance. First we compute the z scores.
x <- sqrt(bikedata$passing.distance)
z <- (x - mean(x))/sd(x)Next, we check what proportion have absolute values less than 1, 2, or 3.
mean(abs(z) <= 1) # Empirical Rule predicts 68%...[1] 0.6994
mean(abs(z) <= 2) # ...95%...[1] 0.9524
mean(abs(z) <= 3) # ...and 99.7%.[1] 0.9936
Using the mean function to compute the proportion of TRUE values looks
unusual the first time you see it, but this is a common idiom in R. It works
by performing the following steps:
- Convert
TRUEandFALSEvalues in the input vector to1and0. - Sum the elements of the converted vector. The sum will be equal to
the number of
TRUEvalues in the original vector. - Divide by the length of the vector. This will give you the proportion
of
TRUEvalues in the original vector.
Integer Indexing
We have already seen slicing (using square brackets). More generally, R supports indexing by any integer vector.
x <- c(-1.1, -1, -3.2, 0.5, 0.9, 0.2, -2.1, 0.1, 0.2, 1)
x[c(3, 1, 7)] # elements 3, 1, and 7[1] -3.2 -1.1 -2.1
Negative indices specify omitted elements:
x[c(-3, -1, -7)] # all elements but 3, 1, and 7[1] -1.0 0.5 0.9 0.2 0.1 0.2 1.0
You are not allowed to mix positive and negative indices:
x[c(-3, 1, 7)] # this is an errorError: only 0's may be mixed with negative subscripts
You are allowed to include 0s in the index set, but doing so has no effect
out the output.
You can use integer indexing in a slightly different form to extract particular rows or columns of a data frame.
bikedata[c(2, 4, 6, 19), ] # rows 2, 4, 6, and 19; all columnsvehicle colour passing.distance street helmet kerb datetime 2 HGV Red 0.998 Urban N 0.5 2006-05-11 16:30:00 4 Car <NA> 1.640 Urban N 0.5 2006-05-11 16:30:00 6 Car Grey 1.509 Urban N 0.5 2006-05-11 16:30:00 19 Car Grey 1.290 Main Y 1.0 2006-05-12 07:46:00 bikelane city 2 N Salisbury 4 N Salisbury 6 N Salisbury 19 N Salisbury
bikedata[c(99, 10, 3), c(2, 4)] # rows 99, 10, 3, 12; columns 2 and 4colour street 99 Grey Main 10 <NA> Main 3 Blue Urban
Application: Top 5 Closest Passes
The next code listing extracts the 5 trips with the shortest passing distances.
shortest <- sort(bikedata$passing.distance, index.return = TRUE)$ix[1:5]
bikedata[shortest, ] vehicle colour passing.distance street helmet kerb
988 Car Blue 0.394 Main N 0.25
2040 LGV White 0.493 Urban N 0.75
1906 Bus Green 0.510 Urban Y 0.50
1590 Car Green 0.527 Main Y 0.75
1043 Car Blue 0.636 Main N 1.25
datetime bikelane city
988 2006-05-20 16:21:00 N Salisbury
2040 2006-06-05 15:02:00 N Bristol
1906 2006-06-05 13:42:00 N Bristol
1590 2006-05-31 10:14:00 N Salisbury
1043 2006-05-26 15:18:00 N Salisbury
The explanation of the code is as follows:
-
First, we call
sortwithindex.return=TRUE. This sorts the values returns a list with two components: adoublevector namedx, which contains the values ofbikedata$passing.distancesorted in increasing order, and an integer vector namedix, which contains the indices of these values. -
We extract the component of the result names
ixwith the$ixoperator. -
Then, we take elements
1:5and store them inshortest. These are the indices of the 5 smallest values, not the values themselves. -
Finally, with the command
bikedata[shortest,], we ask for the rows indicated byshortestand all columns of the data frame.
To get the longest passing distance, we can modify the original code,
adding decreasing=TRUE to the call to sort:
longest <- sort(bikedata$passing.distance, index.return = TRUE, decreasing = TRUE)$ix[1:5]
bikedata[longest, ] vehicle colour passing.distance street helmet kerb
1059 Car Grey 3.787 Urban N 1.25
1946 Car Green 3.571 Urban N 0.25
992 Car Blue 3.560 Main N 0.25
1868 Car Black 3.489 Urban N 0.50
1187 Car Red 3.248 Main N 1.00
datetime bikelane city
1059 2006-05-26 15:18:00 N Salisbury
1946 2006-06-05 14:05:00 N Bristol
992 2006-05-20 16:21:00 N Salisbury
1868 2006-06-05 12:37:00 N Bristol
1187 2006-05-27 09:25:00 N Salisbury
If you want the passing distances to be in ascending order, you can use the
rev function to reverse the indices:
bikedata[rev(longest), ] vehicle colour passing.distance street helmet kerb
1187 Car Red 3.248 Main N 1.00
1868 Car Black 3.489 Urban N 0.50
992 Car Blue 3.560 Main N 0.25
1946 Car Green 3.571 Urban N 0.25
1059 Car Grey 3.787 Urban N 1.25
datetime bikelane city
1187 2006-05-27 09:25:00 N Salisbury
1868 2006-06-05 12:37:00 N Bristol
992 2006-05-20 16:21:00 N Salisbury
1946 2006-06-05 14:05:00 N Bristol
1059 2006-05-26 15:18:00 N Salisbury
Logical Indexing
We can use a logical vector to select particular elements of another vector. To do so, the logical vector should have the same length as the original vector. See the following example:
x <- c(0.5, 1.7, -0.8, -1.4, 0.1, -1, -0.4, -0.8, -0.7, 1.7)
y <- c(0, -1.5, -1, -1, -0.1, 0.7, 1.7, -0.2, 0.1, -0.9)
x[x > 0] # positive values of x[1] 0.5 1.7 0.1 1.7
y[x > 0] # corresponding elements of y[1] 0.0 -1.5 -0.1 -0.9
In the above example, the expression x > 0 evaluates to a logical vector
with the same length as x:
x > 10[1] FALSE FALSE FALSE FALSE FALSE FALSE FALSE FALSE FALSE FALSE
When we use this result as an index vector, we get back all of the elements
where the index vector is TRUE.
Application: Two-Sample Test
Here is a test of difference in means for the two populations: “square root passing distances without helmet” vs. “square root passing distances with helmet”:
x <- sqrt(bikedata$passing.distance)
h <- bikedata$helmet
t.test(x[h == "N"], x[h == "Y"])
Welch Two Sample t-test
data: x[h == "N"] and x[h == "Y"]
t = 5.249, df = 2348, p-value = 1.664e-07
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
0.02039 0.04471
sample estimates:
mean of x mean of y
1.257 1.225
We use logical indexing to extract the sample values drawn from the two populations, then we compare the population means using an unpaired t-test.